- Please read our recent publication for a complete introduction to the dataset:
Robust and accurate prediction of residue-residue interactions across protein interfaces using evolutionary information
Sergey Ovchinnikov, Hetunandan Kamisetty, and David Baker.
Elife (2014). [LINK] [PDF]
|
 |
GREMLIN Results
- E. coli Dataset
- T. thermophilus Dataset
- PDB Dataset (benchmark results)
|
|
GREMLIN Server
Input FASTA Alignments (compressed)
|
| Predicted complexes
|
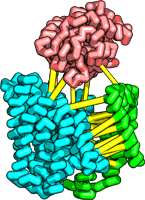
| The TRAP complex |
Tripartite efflux system |
| Download: pdb
|
Download: pdb
|
| Pyruvate formate lyase-activating enzyme complex |
D-methionine transport system |
| Download: pdb
|
Download: pdb
|
Updates
- Clarifications to a few points we bring up in the paper.
- Where is chain "D" in 3A0R (as shown in Figure 3)?
To get chain D you must download the entire PDB (biological assembly), the standard fetch command in PyMol only downloads the asymmeric unit. [A]BC[D] are in the order in which each chain appears in the biological assembly: http://www.rcsb.org/pdb/files/3A0R.pdb1.gz
- Aren't you missing many sequences due to HHblits hard-coded limit of 65535 sequences?
65535 is the largest number that can be held in an unsigned integer. To overcome this, we modified the code slighly as follows:
- hhdecl.C:EXTERN const int MAXSEQ=262140;
- hhfullalignment.C: long unsigned int lq[MAXSEQ];
- hhfullalignment.C: long unsigned int lt[MAXSEQ];
- I am not able to recover the same number of sequences for 3G5O_AB, 1TYG_BA during join.
We somehow managed to omit this part in the final version of the manuscript =[
For the initial E. coli complexes analysis, the same e-value (1E-20) was used throughout. For the PDB benchmark set, given that many of the PDB chains were much shorter than the original E. coli genes and the starting PDB sequence was sometimes very different in identity, this required adjusting our e-value (1E-04) to recover the same number of sequences as in our E. coli alignments. Even though the "e-value" is suppose to be length independent, it tends to break-down when the protein length is less than ~100. For the PDB benchmark set, we used an e-value of 1E-04 for short length proteins.
Please contact us if you have any other questions/concerns!
|
